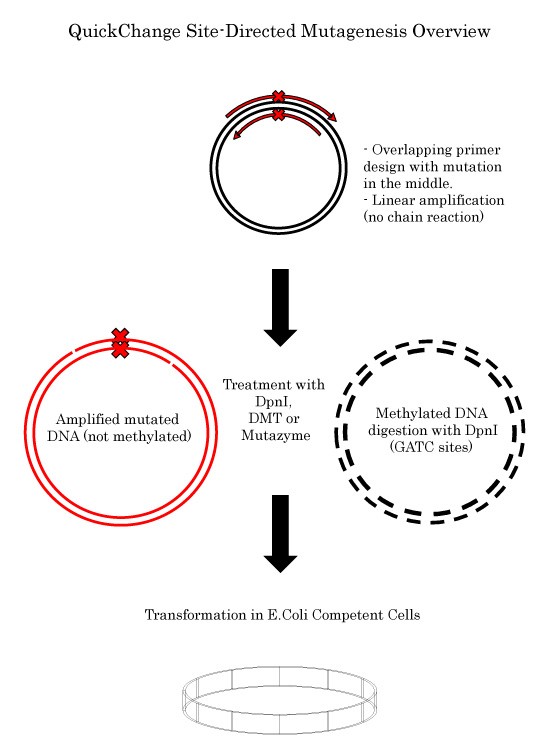
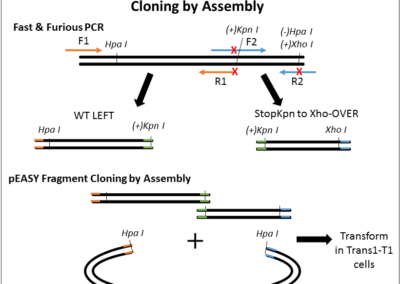
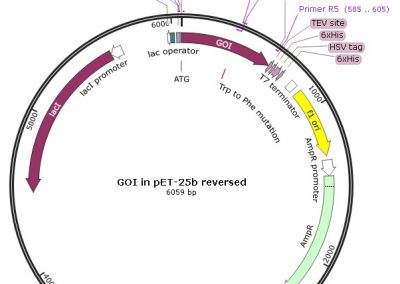
QuickChange Site-Directed Mutagenesis Protocol
Site-Directed Mutagenesis using WVA and complementary primers Go to the Main SDM pageGet inspired by our PCR Success StoriesQuickChange Site-Directed Mutagenesis
Advantages
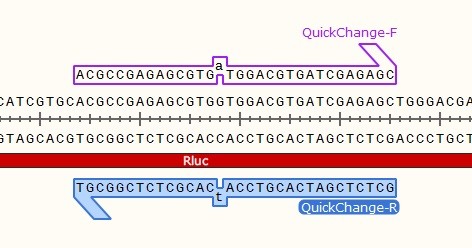
- Both primers contain the mutation.
- In vitro removal of methylated DNA (parental template).
- Simple 3-step workflow***.
Disadvantages
- Long amplification protocol – 25 cycles (between 4 and 8 hours).
- Low yield amplification of DNA.
- Complementary primers anneal together.
- Tandem primer insertion at or near the mutated site often occurs.
- Large insertions are problematic.
- Large deletions are problematic.
***To ensure that no extra mutations are introduced in the vector backbone, site-directed mutagenesis strategies that use whole vector amplification require subcloning of the sequenced insert into a vector that hasn’t been amplified by PCR. Alternatively, the whole vector can be sequenced.
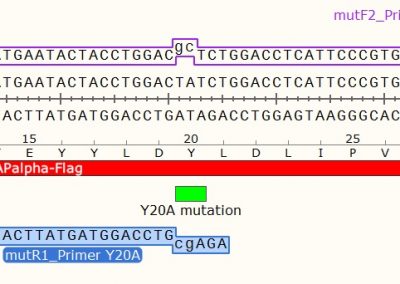
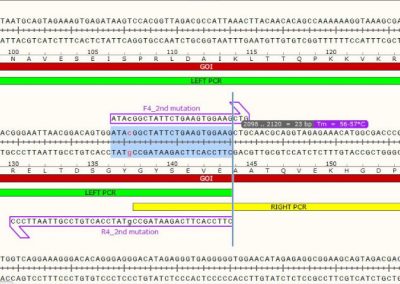
STEP 1 - Primer Design
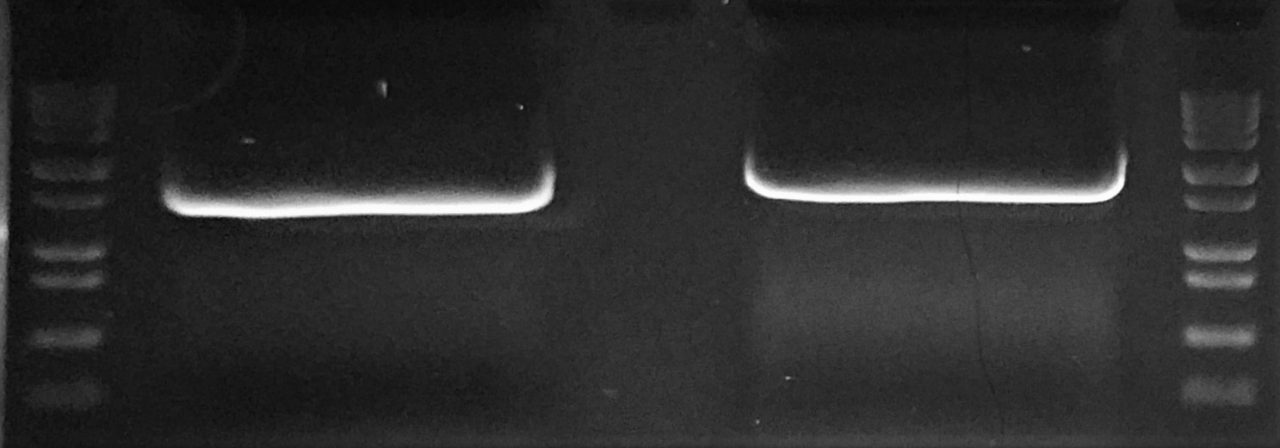
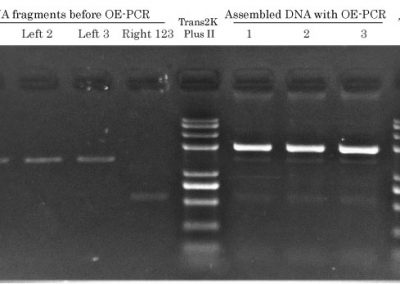
STEP 2 - PCR
QuickChange PCR Protocol
QuickChange PCR Setup
- H2O : to 50 ul
- 5x buffer : 10 ul
- dNTPs (2.5 mM): 4 ul (0.2 mM final)
- Forward Primer (10 uM): 1.25 ul (250 nM final)
- Reverse primer (10 uM): 1.25 ul (250 nM final)
- plasmid DNA : 50 ng
- FastPfu FLY (2.5 u/ul) : 1 ul
QuickChange PCR Cycling
- Denaturation: 120s at 95 °C
- 16-25 x
- Denaturation: 20s at 95 °C
- Annealing: 30s at 55 °C
- Extension: 60 s/kb at 68 °C
- Final extension: 300s at 68 °C
Suggested Products
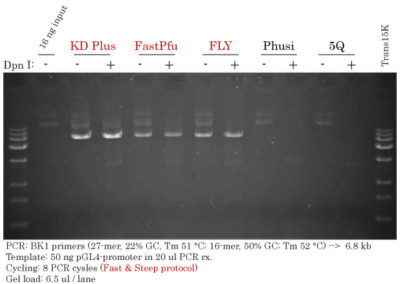
STEP 3 - Digestion
Removal of Methylated Template
Add 1 ul of DMT Enzyme or DpnI and incubate your at 37 °C for 1 h.
STEP 4 - Transformation
Transformation of Competent Cells
- Thaw 50 ul of high-efficiency (> 108 cfu/ug) chemically competent cells on ice.
- Add 0.5 – 5 ul of the QuickChange reaction from Step 3 and gently flick the tube 3 times before incubating on ice for 30 min.
- Heat shock the cells by incubating at precisely 42 °C for 30-45 s (depends on the cells).
- Incubate on ice for > 2 min.
- Add 450 ul of SOC or LB media to the cells, then agitate at 200 rpm, 37 °C for 45-60 min.
- Spread 100-200 ul on a prewarmed LB-agar plate containing the appropriate antibiotic(s).