AT-Rich Vector Amplification by Fast & Steep PCR for Deletion by Site-Directed Mutagenesis
PCR Success Story #20AT-Rich Vector Amplification by Fast & Steep PCR
During an early 2017 customer visit in Quebec city, we met with a researcher who was unsuccessful at performing whole Vector Amplification. The customer was using the Q5® Hoststart Mastermix provided in the Q5® Site-Directed Mutagenesis kit from NEB. At Civic Bioscience, we don’t recommend any site-directed mutagenesis strategies consisting in whole vector amplification (read PCR Success Story #16 for the difference between 2 workflows). However, in this case, primers were already designed by the researcher to be compatible with such strategy (i.e Q5® Site-Directed Mutagenesis) and allow for site-specific deletion of a small 16 bp fragment from a promoter region cloned in a pGL4 backbone vector. One of the foreseable obstacles was the low GC content of the sequence to be amplified. We had to deal with an AT-Rich Vector
To protect the research and privacy of the customer the name of the promoter and primer sequences were slightly modified to be unrecognizable.
Previously, we had developped a new PCR setup and cycling protocol that allows for near 100% integration of all primers in only 5 -10 cycles of PCR.. This new protocol is termed Fast & Steep PCR.
Project Details
Client: Université Laval
Date: January 27th, 2017
Type of experiment: Whole vector amplification AT-Rich PCR, Fast & Steep PCR, Site-Directed Mutagenesis.
DNA Polymerase: TransStart KD Plus, FastPfu and FastPfu FLY High-Fidelity DNA Polymerases
Competitor DNA Polymerases: Phusion® HotStart II and Q5® High-Fidelity DNA Polymerases
Phusion® is a registered trademark of ThermoFisher Scientific. Q5® is a registered trademark of New England Biolabs, Inc.
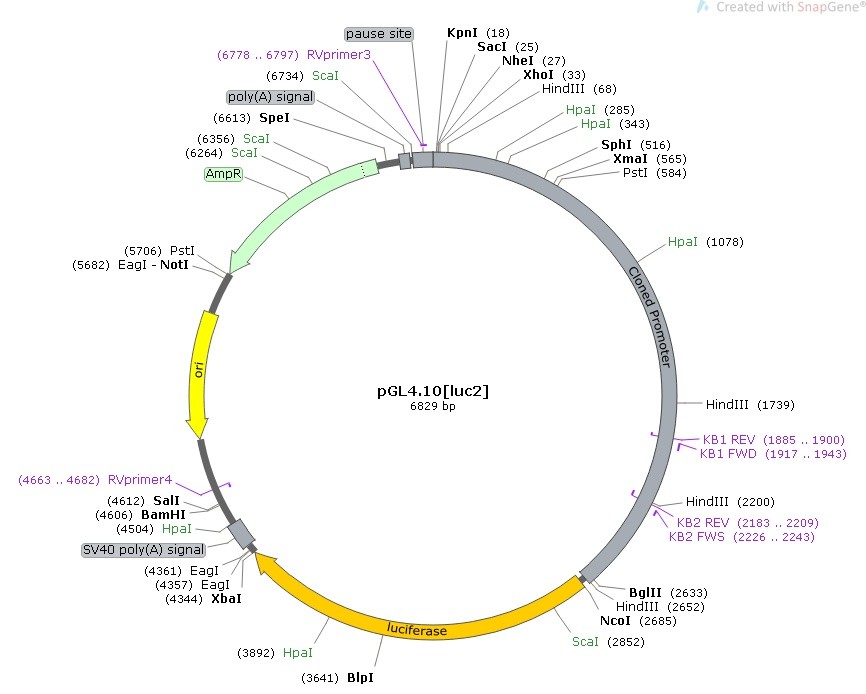
pGL4/promoter map and primer sequences
Template used for AT-Rich Vector Amplification:
- pGL4 vector (Promega) containing the promoter region of interest : 6829 bp (48% GC)
- Insert promoter: 2657 bp (41% GC) – the most AT-Rich region.
Deletion primers used for Vector Amplification #1:
- Forward primer 1: GCACACGCCTGAAAACAT (18-mer; 50% GC; Tm 56 °C)
- Reverse primer 1: CAAATTGCAAAATTGGTTTACTTAGCA (27-mer; 30% GC; Tm 54 56 °C)
Deletion primers used for Vector Amplification #2:
- Forward primer 2: TTTGTATAAAGTTACTTGAGTAAAGTT (27-mer; 22% GC; Tm 51 °C)
- Reverse primer 2: CCAGCCAGCTATAAGT (16-mer; 50% GC; Tm 52 °C)
AT-Rich Vector Amplification #1
Vector Amplification by Fast & Steep PCR
For Vector Amplification #1, KKP (CivicBio), TransStart KD Plus, FastPfu, FastPfu FLY (TransBionovo), Phusion® HotStart II (ThermoFisher) and Q5® (NEB) High-Fidelity DNA Polymerases were compared.
PCR Setup for Phusion and Q5 (undisclosed for other DNA Polymerases Fast & Steep PCR setup)
- H2O : fill to 20 ul
- 5x buffer : 4 ul
- dNTPs (2.5 mM): 1.6 ul (0.2 mM final)
- Forward primer (10 uM): 1 ul (0.5 nM final)
- Reverse primer (10 uM) : 1 ul (0.5 nM final)
- Plasmid : 1 ul (50 ng)
- Phusion or Q5 (2 u/ul) : 0.2 ul
Fast & Steep PCR Cycling for Vector Amplification #1:
- Denaturation: 120s at 95 °C
- 8 x
- Denaturation: 15s at 95 °C
- Annealing: 30s at 51 °C
- Extension: 420s at 68 °C
- Final extension: 600s at 68 °C
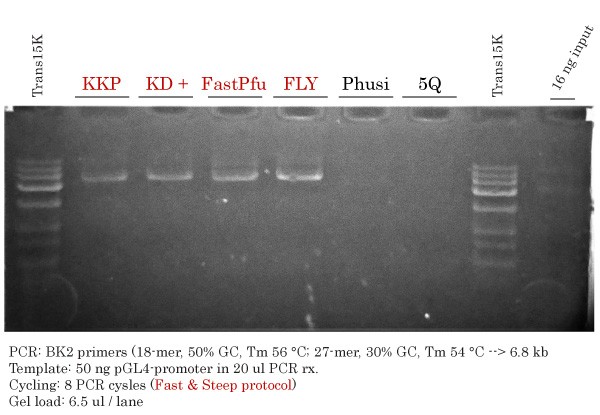
Successful Vector Amplification #1 using TransBionovo DNA Polymerases but not with Phusion or Q5
Whole Vector Amplification using the primer pair #1 was achieved using a low extension temperature (68 °C). KKP, TransStart KD Plus, FastPfu and FastPfu FLY, but not Phusion® HotStart II and Q5® High-Fidelity DNA Polymerases, successfully amplified the intended PCR band at 6.8 kb. The Fast & Steep PCR method implies that a considerable amount of DNA template is introduced in the PCR reaction. The equivalent input that would normally be seen in all samples (16 ng in 6.5 ul out of the 20 ul PCR mix) is shown on the right-hand side. Note that only 8 PCR cycles were performed, thus strongly reducing the chance of introducing undesired mutations during Vector Amplification. FastPfu FLY (Ultra-HiFi) DNA Polymerase was the most specific and efficient.
Trans15K DNA Ladder bands: 15 000, 10 000, 7500, 5000, 3000, 1500, 1000, 500 bp.
AT-Rich Vector Amplification #2
Vector Amplification by Fast & Steep PCR
For Vector Amplification #2, TransStart KD Plus, FastPfu, FastPfu FLY (TransBionovo), Phusion® HotStart II (ThermoFisher) and Q5® (NEB) High-Fidelity DNA Polymerases were compared. Additionnally, we confirmed specific amplification by digesting the methylated template vector using DMT Enzyme (Dpn I).
DpnI cleaves at GATC sites if they are methylated.
PCR Setup for Phusion and Q5 (undisclosed for other DNA Polymerases Fast & Steep PCR setup)
- same as Vector Amplification #1
Fast & Steep PCR Cycling for Vector Amplification #2:
- Denaturation: 120s at 95 °C
- 8 x
- Denaturation: 15s at 95 °C
- Annealing: 20s at 49 °C
- Extension: 420s at 66 °C
- Final extension: 600s at 66 °C
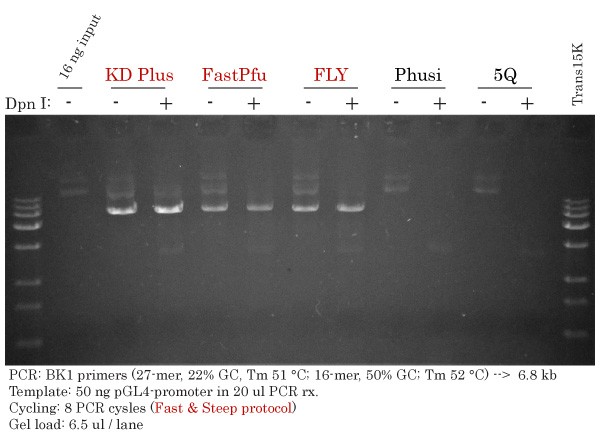
Successful Vector Amplification #2 using TransBionovo DNA Polymerases but not with Phusion or Q5
Whole Vector Amplification using the primer pair #2 was achieved using a low extension temperature (66 °C). Remarkably, TransStart KD Plus, FastPfu and FastPfu FLY, but not Phusion® HotStart II and Q5® High-Fidelity DNA Polymerases, successfully amplified the intended PCR band at 6.8 kb using the Fast & Steep PCR method.
To confirm the nature of the bands observed at or above the 15 kb marker lane, 6.5 ul of PCR products were digested with DMT Enzyme (Dpn I). Indeed, agarose gel electrophoresis (- and + lanes) confirmed these bands to be the input pGL4 template as they were shifted to smaller product sizes in the (+) lanes. In addition, an undigested input template (left-hand side) strongly confirmed this conclusion.
KD Plus (Ultra-HiFi) DNA Polymerase was the most specific and efficient in Vector Amplification #2 using the Fast & Steep PCR protocol.
The Outcome
The difficulty of performing whole Vector Amplification was due to primer and sequence low GC-content – AT-Rich PCR. In contrast to what the researcher experienced using a competitor DNA polymerase, we succeeded at performing BOTH vector amplifications on our very first attempt. However, Vector Amplification # 2 required a simple optimization. In fact, lowering the annealing time to 20 seconds and the extension temperature to 66 °C resulted in better yield. Once more, our results demonstrate that TransBionovo High-Fidelity DNA Polymerases clearly set a Higher Level of DNA Polymerase Performance when compared to the popular Phusion® and Q5® DNA Polymerases.
Now that the PCR reactions are successful, the researcher can proceed to the second step of the Q5® Site-Directed Mutagenesis (KLD incubation).
The new PCR method, named Fast & Steep PCR, shows a very promising alternative to conventional PCR methods such as Site-Directed Mutagenesis.
Fast & Steep PCR enables:
- to perform a very limited amount of PCR cycles on most templates;
- to reduce errors during polymerization;
- the incorporation of desired mutations or extensions to any DNA template;
- to reduce the PCR running time by at least 3-fold.